Arkangel Ai designs an Open Source AI tool for the early detection of Childhood Leukemia for UNICEF

The challenge
According to the latest World Health Organization report, leukemia is one of the three most common childhood cancers worldwide. Due to a lack of experienced staff, health system coverage, and effective screening protocols, leukemia diagnosis is a significant challenge, especially in low- and middle-income countries.
Notably, in Colombia, these challenges in early diagnosis are such that the five-year survival rate for children diagnosed with the disease is estimated at 41%, much lower than the 80% estimated for high-income countries.
We set out to generate an accurate, fast, affordable, and objective solution to diagnose childhood leukemia using Artificial Intelligence; this blog-post is about what we achieved in just 24 hours using the computational power of Hippocrates AutoML.
“Hippocrates AutoML is Arkangel Ai’s new functionality that allows healthcare institutions to use their data resources to generate Ai models according to their medical objectives. Says Leonardo Velásquez Growth Lead at Arkangel Ai”
Project goal
The ai algorithm aims to automatically detect and classify lymphocytes in images of stained blood and bone marrow samples taken with a microscope. The solution has two main components:
- A Deep learning algorithm that identifies and segments the lymphocytes present in the patients’ samples.
- A deep learning classification algorithm that classifies the lymphocytes between normal-lymphocytes and abnormal-lymphocytes (lymphoblastoma).
In this way, if at the end of the patient’s sample analysis, there are more than 20% lymphoblastoma, they will be classified as a leukemia patient.
Model development using Hippocrates, EUREKA!!
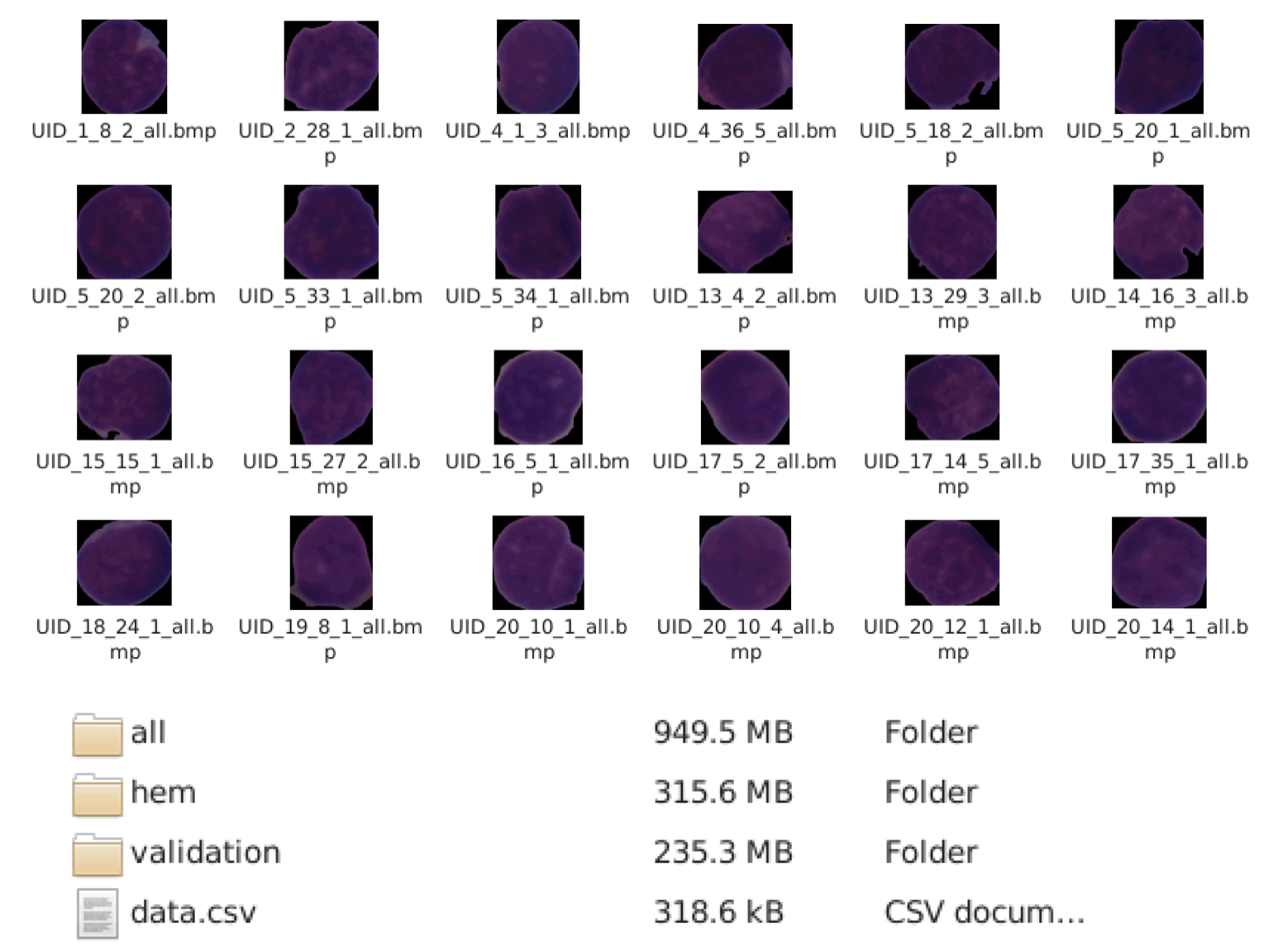
For this project, we use 15.135 images from 118 patients with two labeled classes: “Normal Cell” and “Leukemia Blast.” Once the photos and annotation files were prepared to meet Hippocrates’ requirements, we uploaded them to the cloud platform and we clicked “start experiment,” and Hippocrates started the algorithm construction.
For this real-case experiment, we were very nervous and excited. After 24 hours, first thing in the morning, Hippocrates sent an email notifying us that the experiment was finished successfully, and this is what we found once we logged into the platform:
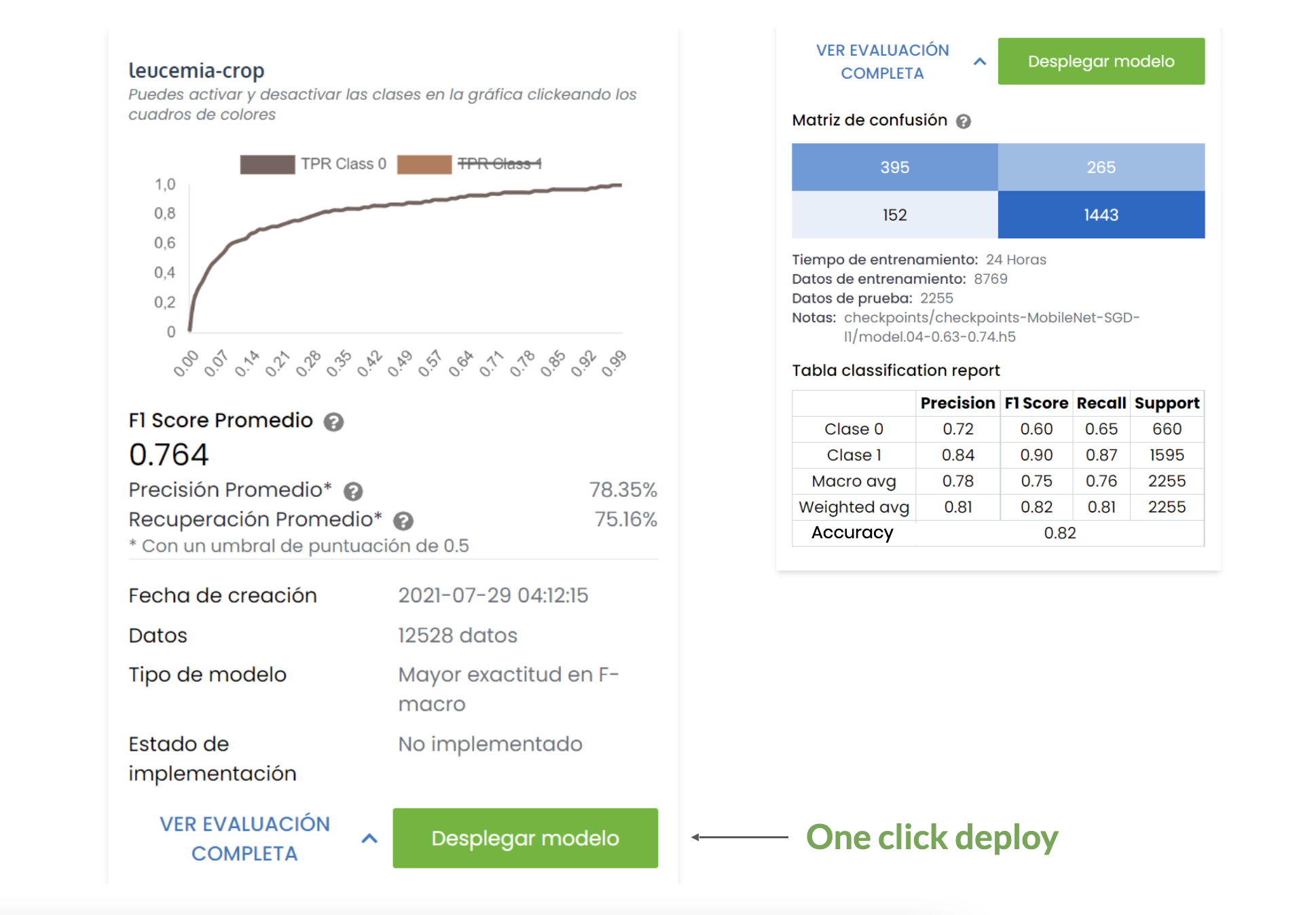
Ranking with the TOP 5 model, the best one had this specifications:
- Accuracy: 0.82
- Sensitivity: 0.91
- Specificity: 0.60
- F1-macro: 0.76
Definitely not bad for a 24-hour experiment!! We celebrated and made some calls to some friends in UNICEF.
Although this first iteration showed great potential and the power of the Hippocrates tool, we are interested in developing a practical algorithm for the detection of leukemia in children. In that sense, our next steps are to improve metrics overall and build the segmentation algorithm. We will also combine both algorithms into a public API to serve as a free service to start piloting in a lab with a partner institution in Colombia.
Project targets/milestones for the next 12 months:
- Implement the prototype in 5 leading Leukemia detection labs in Latam using the open-source API.
- Run 15.000 predictions on patient samples (~1.250/month) across the selected labs. Report the results in precision metrics, cost and time savings, and patient outcomes versus the traditional method.
- Co-create Latam’s first Leukemia Dataset with the help of expert annotators. We will use this dataset to retrain the prototype seeking to improve the metrics previously described. -Close partnerships for improved distribution, repository support, and subsequent funding for two years more. Here are some ideas: Minister of health and social protection, the Colombian Fund for High-Cost Diseases, the Colombian Leukemia and Lymphoma Foundation, and the Colombian National Institute of Health.
Finally, our primary goal is to put Artificial intelligence at the service of the whole medical community. We left here the link to the final model and its future iterations. In case you want to contribute, feel free to open a pull request.





Comments